11. Reporting#
After calculating summary statistics, visualizing them, and testing for significant effects using inferential statistics, we must put these results together to form a narrative that explains what we have found. When reporting results, it is important to describe the results thoroughly and accurately, and in a way that is understandable to a relatively wide audience. It is also important to publish your code with documentation, to allow others to reproduce your results.
When completing exercises, you may find it helpful to have the chapter website and cheat sheet open in a browser.
There are many different kinds of analyses that we may want to explore when analyzing a dataset. We will use examples from analysis of the Morton et al. (2013) dataset, to illustrate how to run and report different analyses. We will use Polars, Seaborn, and Pingouin to get the plots and statistics we need to make conclusions about the dataset.
First, we will load the data and filter out invalid recall attempts.
import polars as pl
import seaborn as sns
import pingouin as pg
from datascipsych import datasets
data_file = datasets.get_dataset_file("Morton2013")
raw = pl.read_csv(data_file, null_values="n/a")
data = raw.filter(pl.col("study")) # remove invalid recall attempts
data.head()
| subject | session | list | item | input | output | study | recall | repeat | intrusion | list_type | list_category | category | response | response_time |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| i64 | i64 | i64 | str | i64 | i64 | bool | bool | i64 | bool | str | str | str | i64 | f64 |
| 1 | 1 | 1 | "TOWEL" | 1 | 13 | true | true | 0 | false | "pure" | "obj" | "obj" | 3 | 1.517 |
| 1 | 1 | 1 | "LADLE" | 2 | null | true | false | 0 | false | "pure" | "obj" | "obj" | 3 | 1.404 |
| 1 | 1 | 1 | "THERMOS" | 3 | null | true | false | 0 | false | "pure" | "obj" | "obj" | 3 | 0.911 |
| 1 | 1 | 1 | "LEGO" | 4 | 18 | true | true | 0 | false | "pure" | "obj" | "obj" | 3 | 0.883 |
| 1 | 1 | 1 | "BACKPACK" | 5 | 10 | true | true | 0 | false | "pure" | "obj" | "obj" | 3 | 0.819 |
In this study, participants studied lists of 24 items, which were celebrities, famous locations, or common objects. After each list, participants were asked to freely recall the items from that list in whatever order they wanted. In the dataset, trials where the recall column is true indicate that the participant did successfully recall that item. There were two types of list: pure lists (all the same category) and mixed lists (drawn from different categories).
11.1. Summary statistics#
When reporting how a dependent variable varies depending on an independent variable, it’s important to report summary statistics for each value of the independent variable, including the mean and standard error of the mean (SEM).
For example, say we are analyzing the Morton et al. (2013) dataset and want to know whether recall varies by stimulus category (celebrity, location, or object).
First, we need to calculate the mean of the dependent variable (here, recall) for each subject and value of the independent variable (here, category). We can do this using group_by with agg.
cat = (
data.group_by("subject", "category")
.agg(pl.col("recall").mean())
.sort("subject", "category")
)
cat.head(6)
| subject | category | recall |
|---|---|---|
| i64 | str | f64 |
| 1 | "cel" | 0.6015625 |
| 1 | "loc" | 0.481771 |
| 1 | "obj" | 0.4453125 |
| 2 | "cel" | 0.653646 |
| 2 | "loc" | 0.513021 |
| 2 | "obj" | 0.489583 |
When writing about the effect of stimulus category on recall, we should report the mean recall across subjects for each category. We can calculate the mean using group_by and agg again, this time to average across subjects.
(
cat.group_by("category")
.agg(pl.col("recall").mean())
.sort("category")
)
| category | recall |
|---|---|
| str | f64 |
| "cel" | 0.548372 |
| "loc" | 0.517383 |
| "obj" | 0.483984 |
Looking at the mean recall for each category, we can see that celebrities were best recalled (mean recall: 0.55), followed by locations (mean recall: 0.52), and then objects (mean recall: 0.48).
When reporting the mean values of some dependent variable, it’s best practice to also report the standard error of the mean (SEM), which indicates uncertainty in the estimate of the mean. The SEM gives a sense of how reliable our estimate of the mean is. The SEM is defined as:
To calculate the SEM, we need to get the standard deviation of our dependent variable and divide it by the square root of the number of samples (here, the number of subjects).
We can use Polars to calculate the SEM using an expression. We take the column with our dependent measure and calculate the standard deviation \(\sigma\) using the .std function. Then we divide that by \(\sqrt{n}\), using the .len function to get the number of subjects, followed by .sqrt to take the square root of that.
pl.col("recall").std() / pl.col("recall").len().sqrt()
Like we did for calculating the mean, we can use this expression with group_by and agg to calculate the SEM for each condition.
(
cat.group_by("category")
.agg(pl.col("recall").std() / pl.col("recall").len().sqrt())
.sort("category")
)
| category | recall |
|---|---|
| str | f64 |
| "cel" | 0.013505 |
| "loc" | 0.015963 |
| "obj" | 0.018889 |
Note that, for a given standard deviation \(\sigma\), increasing the number of samples \(n\) will decrease the SEM. The more samples we have, the more confident we can be in our estimate of the mean. Lower SEM indicates less error, or greater confidence.
We can use Polars to calculate the mean and SEM together in one expression.
(
cat.group_by("category")
.agg(
mean=pl.col("recall").mean(),
sem=pl.col("recall").std() / pl.col("recall").len().sqrt(),
)
.sort("category")
)
| category | mean | sem |
|---|---|---|
| str | f64 | f64 |
| "cel" | 0.548372 | 0.013505 |
| "loc" | 0.517383 | 0.015963 |
| "obj" | 0.483984 | 0.018889 |
We can then use this information to summarize our results when writing about them. For example, here we could say:
Recall varied by stimulus category, with the greatest recall for celebrities (mean: 0.548, SEM: 0.014), then locations (mean: 0.517, SEM: 0.016), and least recall for objects (mean: 0.484, SEM: 0.019).
Next, we will demonstrate a series of analysis types using this dataset. In each case, reporting results of an analysis involves visualizing an effect of a variable (or lack of effect), summarizing the effect using summary statistics, and testing to see if the effect is statistically significant.
11.2. Comparing two conditions#
If we have some dependent variable that was measured in two different conditions, we may want to know if that variable changes reliably depending on the condition.
In this example, we will analyze whether recall accuracy on pure lists is different from recall accuracy on mixed lists.
Calculating measures#
We are interested in how the dependent variable, recall, varies with the independent variable of list type (mixed or pure).
We first need to calculate the mean for each condition, separately for each subject.
lt = (
data.group_by("subject", "list_type")
.agg(pl.col("recall").mean())
.sort("subject", "list_type")
)
lt.head()
| subject | list_type | recall |
|---|---|---|
| i64 | str | f64 |
| 1 | "mixed" | 0.476389 |
| 1 | "pure" | 0.564815 |
| 2 | "mixed" | 0.55 |
| 2 | "pure" | 0.555556 |
| 3 | "mixed" | 0.484722 |
Now we have the mean recall for each list type, separately for each subject.
Visualization#
Next, we can visualize the means and uncertainty in the means using a bar plot with error bars. Seaborn will automatically calculate the means across subjects, and use a bootstrap method to determine error bars around the means.
(
sns.catplot(data=lt, x="list_type", y="recall", kind="bar", height=2)
.set_axis_labels("List type", "Recall probability")
);
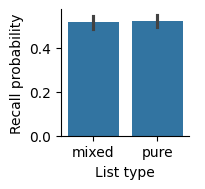
By default, Seaborn will label axes with the names of the columns we used, but those names aren’t always the best for making a visualization; for example, they may have things like underscores in the names. We can use the set_axis_labels method to change the x-axis and y-axis labels to something that is easier to read.
In our report, we should include a figure caption to explain what’s in the figure.
Recall, measured as the fraction of items recalled in each list, was similar for pure and mixed lists. Error bars indicate 95% bootstrap confidence intervals.
Summary statistics#
In addition to having a plot showing the means, it’s helpful to include the mean and SEM of each condition in the text of a report also.
We will use another pair of group_by and agg calls to get the mean and SEM for each list type.
(
lt.group_by("list_type")
.agg(
mean=pl.col("recall").mean(),
sem=pl.col("recall").std() / pl.col("recall").len().sqrt(),
)
.sort("list_type")
)
| list_type | mean | sem |
|---|---|---|
| str | f64 | f64 |
| "mixed" | 0.514097 | 0.014944 |
| "pure" | 0.520718 | 0.013915 |
Inferential statistics#
Finally, we use a t-test to see if there is a significant difference between conditions. We can use a pivot table to access recall for the mixed and pure lists separately.
p = lt.pivot("list_type", index="subject", values="recall")
pg.ttest(p["mixed"], p["pure"], paired=True)
| T | dof | alternative | p_val | CI95 | cohen_d | power | BF10 | |
|---|---|---|---|---|---|---|---|---|
| T_test | -0.997508 | 39 | two-sided | 0.324667 | [-0.02, 0.01] | 0.072498 | 0.073222 | 0.271 |
Because list type is a within-subjects condition, we use a paired t-test. Because \(p>0.05\), we conclude that there is not a significant effect of list type on recall.
Results#
We can report our findings using the figure and something like the following text.
We examined whether there was a difference in recall between the different list types. We found that there was similar recall in the mixed (mean=0.514, SEM=0.081) and pure (mean=0.521, SEM=0.082) lists. There was not a significant difference in recall based on list type (t(39)=1.00, p=0.33, Cohen’s d=0.072).
When reporting statistics, we give the mean and SEM for each condition, while providing text to help explain the results. We then report the results of the hypothesis test, giving the statistic, the degrees of freedom, the p-value, and a measure of effect size (here, Cohen’s d is commonly used for t-tests).
The sign of the t-statistic is usually discarded. It depends on whether we test (mixed - pure) or (pure - mixed), but we just care about whether there is a difference. The reader can tell the direction of the (non-significant) effect by looking at the means, rather than the sign of the t-statistic.
11.3. Comparing multiple conditions#
If we have more than two conditions, we can use a one-way ANOVA to test for differences. In this example, we will examine whether recall varies depending on the serial position of an item. We will split up items by whether they were studied in an early position in the list (1-8), a middle position (9-16), or a late position (17-24).
Calculating measures#
First, we will have to add a new column that indicates serial position group. We can use the cut method, which divides a column into bins based on the boundaries we indicate. We can provide a label for each bin.
sp = data.with_columns(
serial_position=pl.col("input").cut([8.5, 16.5], labels=["early", "middle", "late"])
)
sp.head(10)
| subject | session | list | item | input | output | study | recall | repeat | intrusion | list_type | list_category | category | response | response_time | serial_position |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| i64 | i64 | i64 | str | i64 | i64 | bool | bool | i64 | bool | str | str | str | i64 | f64 | cat |
| 1 | 1 | 1 | "TOWEL" | 1 | 13 | true | true | 0 | false | "pure" | "obj" | "obj" | 3 | 1.517 | "early" |
| 1 | 1 | 1 | "LADLE" | 2 | null | true | false | 0 | false | "pure" | "obj" | "obj" | 3 | 1.404 | "early" |
| 1 | 1 | 1 | "THERMOS" | 3 | null | true | false | 0 | false | "pure" | "obj" | "obj" | 3 | 0.911 | "early" |
| 1 | 1 | 1 | "LEGO" | 4 | 18 | true | true | 0 | false | "pure" | "obj" | "obj" | 3 | 0.883 | "early" |
| 1 | 1 | 1 | "BACKPACK" | 5 | 10 | true | true | 0 | false | "pure" | "obj" | "obj" | 3 | 0.819 | "early" |
| 1 | 1 | 1 | "JACKHAMMER" | 6 | 7 | true | true | 0 | false | "pure" | "obj" | "obj" | 1 | 1.212 | "early" |
| 1 | 1 | 1 | "LANTERN" | 7 | null | true | false | 0 | false | "pure" | "obj" | "obj" | 2 | 0.888 | "early" |
| 1 | 1 | 1 | "DOORKNOB" | 8 | 11 | true | true | 0 | false | "pure" | "obj" | "obj" | 3 | 0.915 | "early" |
| 1 | 1 | 1 | "SHOVEL" | 9 | 9 | true | true | 0 | false | "pure" | "obj" | "obj" | 1 | 1.287 | "middle" |
| 1 | 1 | 1 | "WATER GUN" | 10 | null | true | false | 0 | false | "pure" | "obj" | "obj" | 1 | 0.839 | "middle" |
Next, we calculate the mean for each subject and serial position group.
recency = (
sp.group_by("subject", "serial_position")
.agg(pl.col("recall").mean())
.sort("subject", "serial_position")
)
Visualization#
Now we can visualize the effect of serial position on recall. Because serial position is an ordered variable, we’ll use a line plot, which connects data points instead of a bar plot, which treats conditions as having no particular order.
# hack to get around data conversion problems with categorical data
recency_pd = recency.to_pandas()
recency_pd["serial_position"] = recency_pd["serial_position"].cat.set_categories(["early", "middle", "late"])
(
sns.relplot(data=recency_pd, x="serial_position", y="recall", kind="line", height=2)
.set(ylim=(0, 1), xlabel="Serial position", ylabel="Recall probability")
);
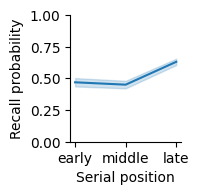
Recall probability varied with serial position group, with the greatest recall in late serial positions. Error band indicates 95% bootstrap confidence intervals.
Summary statistics#
We also calculate summary statistics, including the mean and SEM.
(
recency.group_by("serial_position")
.agg(
mean=pl.col("recall").mean(),
sem=pl.col("recall").std() / pl.col("recall").len().sqrt(),
)
)
| serial_position | mean | sem |
|---|---|---|
| cat | f64 | f64 |
| "early" | 0.468945 | 0.017261 |
| "middle" | 0.45026 | 0.015238 |
| "late" | 0.630534 | 0.013084 |
Inferential statistics#
Finally, we test whether recall varies by serial position group. Because we have more than two groups, we have to use an ANOVA instead of a t-test. Because serial position is a within-subject factor, we use a repeated-measures ANOVA.
pg.rm_anova(data=recency.to_pandas(), dv="recall", within="serial_position", subject="subject")
| Source | ddof1 | ddof2 | F | p_unc | p_GG_corr | ng2 | eps | sphericity | W_spher | p_spher | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | serial_position | 2 | 78 | 205.232427 | 8.456487e-32 | 6.016369e-24 | 0.418096 | 0.732216 | False | 0.634283 | 0.000175 |
The p-value of 1.555e-35 uses scientific notation. In scientific notation, “e” followed by a number is a power of 10. For example, 1.2e2 would be \(1.2x10^2=1.2x100=1200\). Scientific notation is usually used for very large or very small numbers. In this case, there is a negative exponent, so here we have a very small number, \(p=1.56x10^{-35}\). Because \(p<0.05\), we conclude that there is a significant effect of serial position on recall.
Results#
We can report our results with text like this:
We examined whether there was a difference in recall based on serial position group (early positions: 1-8; middle: 9-16; late: 17-24). Late serial positions were recalled best (mean=0.655, SEM=0.104), followed by early serial positions (mean=0.466, SEM=0.074), with the worst recall for middle positions (mean=0.452, SEM=0.071). A one-way repeated measures ANOVA found a significant effect of serial position group (F(2,78)=265.5, p=1.6x10^-35, ng2=0.49).
This follows the same basic pattern as reporting a comparison of two conditions, but now we report the results of the ANOVA. The ANOVA has a different measure of effect size called general eta squared, which measures the size of the effect of serial position group on recall.
11.4. Testing for an interaction between conditions#
Sometimes, the effect of one variable will depend on another variable. For example, someone’s height probably will not usually correlate with their income. But if you split people up by whether or not they are professional basketball players, you might find an interaction, where height predicts income for professional basketball players, but not for non-basketball players.
Calculating measures#
We will examine whether the relationship between category and recall depends on whether the list was pure (all the same category) or mixed (including 8 items from each of the 3 categories). First, we calculate the mean recall for each combination of conditions.
lt_cat = (
data.group_by("subject", "list_type", "category")
.agg(pl.col("recall").mean())
.sort("subject", "list_type", "category")
)
lt_cat.head(6)
| subject | list_type | category | recall |
|---|---|---|---|
| i64 | str | str | f64 |
| 1 | "mixed" | "cel" | 0.5875 |
| 1 | "mixed" | "loc" | 0.433333 |
| 1 | "mixed" | "obj" | 0.408333 |
| 1 | "pure" | "cel" | 0.625 |
| 1 | "pure" | "loc" | 0.5625 |
| 1 | "pure" | "obj" | 0.506944 |
Visualization#
We can visualize how recall varies with list type and category using a bar plot that uses a different hue for each category.
(
sns.catplot(lt_cat, x="list_type", y="recall", hue="category", kind="bar", palette="colorblind", height=2)
.set_axis_labels("List type", "Recall probability")
);
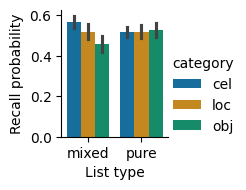
Recall probability by list type and category. Recall was similar for the different categories in the pure lists, but differed between categories in the mixed lists. Error bars indicate 95% bootstrap confidence intervals.
Summary statistics#
We also calculate summary statistics, including mean and SEM.
(
lt_cat.group_by("list_type", "category")
.agg(
mean=pl.col("recall").mean(),
sem=pl.col("recall").std() / pl.col("recall").len().sqrt(),
)
.sort("list_type", "category")
)
| list_type | category | mean | sem |
|---|---|---|---|
| str | str | f64 | f64 |
| "mixed" | "cel" | 0.567604 | 0.014506 |
| "mixed" | "loc" | 0.516771 | 0.01691 |
| "mixed" | "obj" | 0.457917 | 0.020252 |
| "pure" | "cel" | 0.516319 | 0.013552 |
| "pure" | "loc" | 0.518403 | 0.016545 |
| "pure" | "obj" | 0.527431 | 0.017991 |
Inferential statistics#
To test whether list type and category, or some combination of list type and category, affects recall, we can use a two-way ANOVA. If there is a significant list type x category interaction, that means that the effect of category depends on list type (and vice-versa).
Because the list type and category variables are within-subject variables, we calculate a repeated-measures ANOVA using the rm_anova function from Pingouin. This function does not yet support Polars DataFrames, so we must first convert to Pandas format using to_pandas.
df = lt_cat.to_pandas()
pg.rm_anova(df, dv="recall", within=["list_type", "category"], subject="subject")
| Source | SS | ddof1 | ddof2 | MS | F | p_unc | p_GG_corr | ng2 | eps | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | list_type | 0.002630 | 1 | 39 | 0.002630 | 0.995022 | 3.246672e-01 | 3.246672e-01 | 0.000998 | 1.000000 |
| 1 | category | 0.097177 | 2 | 78 | 0.048588 | 6.690909 | 2.080452e-03 | 3.945953e-03 | 0.035599 | 0.828202 |
| 2 | list_type * category | 0.146670 | 2 | 78 | 0.073335 | 49.788702 | 1.162971e-14 | 2.748265e-14 | 0.052772 | 0.969714 |
These results show that there is not a significant main effect of list type. There is a main effect of category. As we can tell from the bar plot, there is a large interaction effect, where category affects recall on mixed lists much more than on pure lists.
Results#
We can report our results using text like this:
We examined whether recall varies based on list type and category using a two-way repeated-measures ANOVA with a Greenhouse-Geisser correction for non-sphericity. We found that recall varied between categories on the mixed lists, with the best recall for celebrities (mean=0.568, SEM=0.015), followed by locations (mean=0.517, SEM=0.017), then objects (mean=0.458, SEM=0.020). On pure lists, recall was similar for the different categories, with the greatest recall for objects (mean=0.527, SEM=0.018), followed by locations (mean=0.518, SEM=0.017), then celebrities (mean=0.516, SEM=0.014). There was no main effect of list type (F(1,39)=1.00, p=0.33, ng2=0.001). There was a main effect of category (F(2,78)=6.69, p=0.004, ng2=0.036). There was also a significant interaction between list type and category (F(2,78)=49.8, p=2.8x10^-14, ng2=0.053).
The Greenhouse-Geisser correction is used to avoid an assumption of standard ANOVAs that different conditions will be equally correlated with one another. In this case, the three categories may not be equally correlated, and the p-GG-corr column has a p-value that is corrected for this possibility.
11.5. Correlating measures#
Sometimes we have two measures that may or may not be related, and we want to examine how they co-vary with each other. For example, we might want to know whether differences in education tend to correlate with differences in income, or whether short-term memory capacity is related to IQ. We cannot use statistics to determine whether two variables are causally related, but we can check to see if they are at least correlated.
Calculating measures#
During the study period, participants had to make a judgment about each item when it appeared. They were asked to make a different rating depending on whether the item was a celebrity (“how much do you love or hate this person?”), a famous landmark (“how much would you like to visit this place?”), or an object (“how often do you encounter this object in your daily life?”). Say that we think participants who take longer to make responses might be processing the items more deeply, and therefore end up having better memory for those items. If our hypothesis is correct, then response time should be correlated with recall.
To test our hypothesis, let’s get the mean recall and mean response time for each subject.
indiv = (
data.group_by("subject")
.agg(
pl.col("recall").mean(),
pl.col("response_time").mean(),
)
.sort("subject")
)
indiv.head()
| subject | recall | response_time |
|---|---|---|
| i64 | f64 | f64 |
| 1 | 0.509549 | 0.792333 |
| 2 | 0.552083 | 1.659799 |
| 3 | 0.480903 | 1.536678 |
| 4 | 0.547743 | 1.67389 |
| 5 | 0.6484375 | 2.307194 |
Visualization#
We can use a scatter plot to visualize the relationship between response time and recall, along with a fit to a linear model, using lmplot.
(
sns.lmplot(indiv, x="response_time", y="recall", height=2)
.set_axis_labels("Response time (s)", "Recall probability")
);
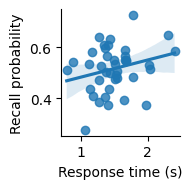
Scatter plot showing the relationship between response time (in seconds) and recall probability across subjects. Line indicates best-fitting linear regression; confidence band indicates a 95% bootstrap confidence interval.
It seems like there might be a slight positive correlation. But is it real, or just due to chance?
Inferential statistics#
We can calculate a correlation to measure the strength of relationship between two variables. The p-value will estimate the probability of having observed a relationship at least this strong due to chance.
pg.corr(indiv["response_time"], indiv["recall"])
| n | r | CI95 | p_val | BF10 | power | |
|---|---|---|---|---|---|---|
| pearson | 40 | 0.253136 | [-0.06, 0.52] | 0.115026 | 0.653 | 0.356031 |
We find that p is greater than 0.05 and conclude that we cannot reject the null hypothesis that the relationship is due to chance.
Results#
We can use the following text to report our results:
We examined whether response time on the encoding task was related to subsequent recall probability. We found that there was not a significant correlation (r=0.253, p=0.12).
11.6. Using Markdown formatting#
GitHub projects generally include a README.md file with information about the project. These files can use special formatting characters to indicate how text should be displayed.
Markdown files can be viewed either as raw Markdown code, or as a rendered display that is nicely formatted for reading. Open the README.md file in the datascipsych project. In Visual Studio Code, you can open a rendered version of the file by clicking on the icon in the top right with a magnifying glass over two boxes. Notice how headings (lines that start with #, ##, or ###) are displayed as headings, sub-headings, or sub-sub-headings. It’s also possible to include links and code blocks. See the Markdown Cheat Sheet for more information.
If you use a GitHub project to track your code, the GitHub website will automatically display a rendered version of your README on the main page of your project. See the datascipsych page for an example.
Installation instructions#
Analysis project README files should include instructions for installing any necessary code. Code formatting can be used to make it easier to read code snippets in your instructions.
Installation instructions depend on how the user is running your code. If you assume they are using Visual Studio Code (which is fine for your final project), you can use instructions like this:
Open the Terminal (
View > Terminal) and runuv sync.
Double-click on the cell above to see what the Markdown code looks like. In this example, the text snippet is placed in a quote box to separate it from the other text. You can use a quote box by placing a > at the start of a line.
You can use backticks (``) to include code in your Markdown file; this code will be formatted with a fixed-with font to differentiate it from regular text.
Run instructions#
Code formatting can also make it easier to read directions for running code, by making filenames and commands stand out. For example, instructions on running analysis code could look like this:
To run the project analysis code, click on
jupyter/project.ipynbin the Explorer pane to open the notebook. You should see a button on the upper right that saysSelect Kernel. Click it to select a kernel to run the notebook. SelectPython Environments, then select the virtual environment created byuv. Finally, clickRun Allto run all cells in the notebook.
Code blocks#
Sometimes, it is useful to include longer code snippets in a README file (for example, to show instructions on how to use a function in one of your code modules). You can include code blocks by surrounding them with triple backticks (```). For example, look at the source code below:
def add_numbers(a, b):
return a + b
For code blocks, you can include a language tag (for example, in the above code, python is placed after the opening backticks) to indicate how the code should be rendered. When there is a language tag, rendered code will include syntax highlighting, which automatically identifies different parts of the code, such as keywords, function names, and variables, and displays them in different colors.
11.7. Including datasets in a project#
For convenience, dataset files may be included with a code project.
A simple approach to including a dataset is to have it in the same directory as any notebooks that are used to analyze it. This is the approach taken in the course project (see the project directory in the datascipsych repository).
A more flexible approach is to include the file within a Python package. This is the approach used by the datascipsych package, which includes multiple datasets that can be analyzed from any directory as long as the datascipsych package is installed.
Datasets in the datascipsych project#
When using code in the datascipsych project, it is not necessary for users to download datasets, because dataset files are included in the source code. See the src/datascipsych/data directory, which includes CSV files for the Morton2013, Onton2005, and Osth2019 datasets.
After installing the datascipsych project, you can get the path to a CSV file using the datasets module. For example, to load the Morton 2013 free-recall dataset:
import polars as pl
from datascipsych import datasets
dataset_file = datasets.get_dataset_file("Morton2013")
data = pl.read_csv(dataset_file)
Look at the src/datascipsych/datasets.py file to see the source code for the get_dataset_file function. It uses the importlib.resources module to determine where the dataset file is installed.
11.8. Summary#
Writing and reporting analyses#
We demonstrated different kinds of common analyses and how they should be reported.
t-test : compare two conditions
One-way ANOVA : compare three or more conditions
Two-way ANOVA : test for an interaction between two variables
Correlation : test whether two variables co-vary
Each type of analysis involves calculating the measure(s) of interest, visualizing the results, calculating summary statistics, and calculating inferential statistics. Finally, this information is put together to write a report of the results.
Using Markdown formatting#
Headers and code formatting can make it easier to read Markdown files such as the README.md file for a project.
11.9. Datasets#
Datasets can optionally be included with an analysis project to make analysis more convenient. If the dataset is not included directly in the project, write information about downloading it to the README.md file.
